Input interpretation
degradation of pyrimidine deoxyribonucleosides (human metabolic pathway)
Pathway topology
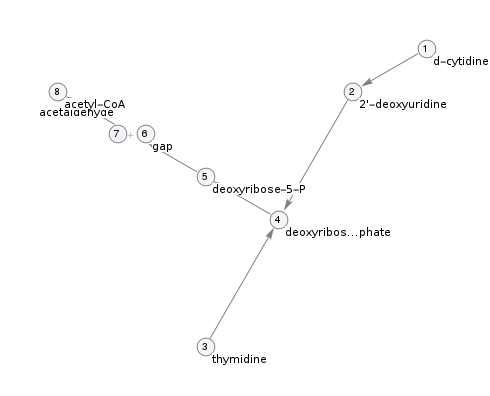
Pathway topology
Pathway components
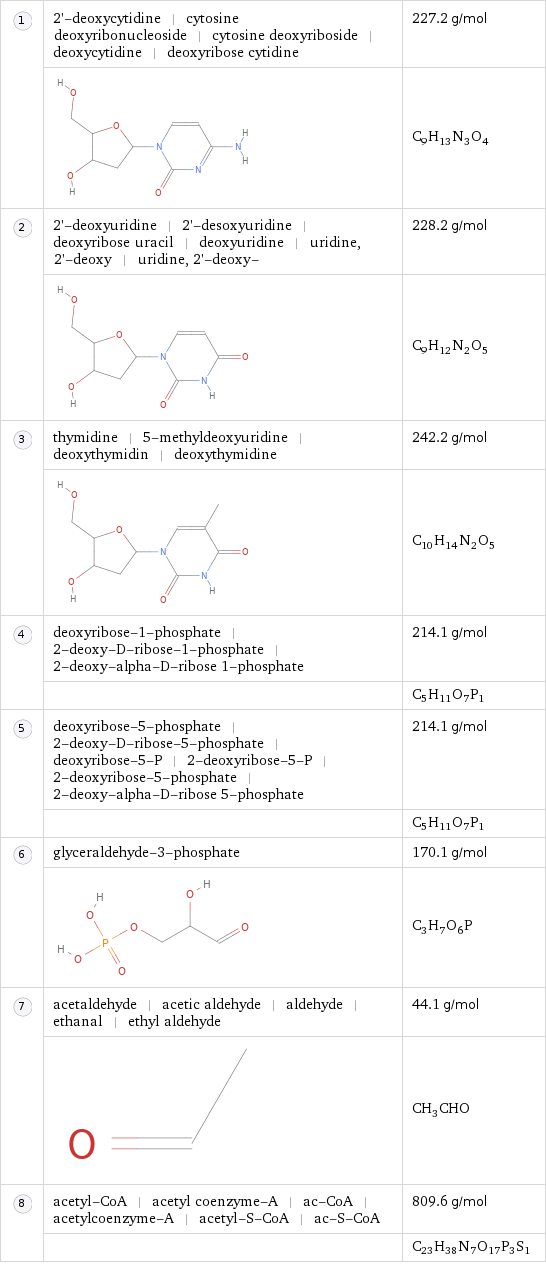
| 2'-deoxycytidine | cytosine deoxyribonucleoside | cytosine deoxyriboside | deoxycytidine | deoxyribose cytidine | 227.2 g/mol | | C_9H_13N_3O_4 | 2'-deoxyuridine | 2'-desoxyuridine | deoxyribose uracil | deoxyuridine | uridine, 2'-deoxy | uridine, 2'-deoxy- | 228.2 g/mol | | C_9H_12N_2O_5 | thymidine | 5-methyldeoxyuridine | deoxythymidin | deoxythymidine | 242.2 g/mol | | C_10H_14N_2O_5 | deoxyribose-1-phosphate | 2-deoxy-D-ribose-1-phosphate | 2-deoxy-alpha-D-ribose 1-phosphate | 214.1 g/mol | | C_5H_11O_7P_1 | deoxyribose-5-phosphate | 2-deoxy-D-ribose-5-phosphate | deoxyribose-5-P | 2-deoxyribose-5-P | 2-deoxyribose-5-phosphate | 2-deoxy-alpha-D-ribose 5-phosphate | 214.1 g/mol | | C_5H_11O_7P_1 | glyceraldehyde-3-phosphate | 170.1 g/mol | | C_3H_7O_6P | acetaldehyde | acetic aldehyde | aldehyde | ethanal | ethyl aldehyde | 44.1 g/mol | | CH_3CHO | acetyl-CoA | acetyl coenzyme-A | ac-CoA | acetylcoenzyme-A | acetyl-S-CoA | ac-S-CoA | 809.6 g/mol | | C_23H_38N_7O_17P_3S_1
Pathway steps

| reaction ⟶ | + H_2O ⟶ + ammonia | activation-induced cytidine deaminase | cytidine deaminase ⟶ | + phosphate ⟶ + uracil | (none) ⟶ | + phosphate ⟶ + thymine | thymidine phosphorylase precursor ⟶ | ⟶ | (none) ⟶ + | ⟶ + | putative deoxyribose-phosphate aldolase ⟶ | + coenzyme A + NAD+ ⟶ + NADH | (none)
Related pathways

| overlapping components (deoxy)ribose phosphate degradation | purine deoxyribonucleosides degradation | mixed acid fermentation | salvage pathways of purine and pyrimidine nucleotides | TCA cycle | glutaryl-CoA degradation |