Input interpretation
adenosine
Chemical names and formulas

formula | C_10H_13N_5O_4 name | adenosine IUPAC name | (3R, 4S, 5R)-2-(6-aminopurin-9-yl)-5-(hydroxymethyl)oxolane-3, 4-diol alternate names | adenocard | adenoscan | beta-adenosine | boniton | myocol | sandesin mass fractions | C (carbon) 44.9% | H (hydrogen) 4.9% | N (nitrogen) 26.2% | O (oxygen) 23.9%
Lewis structure
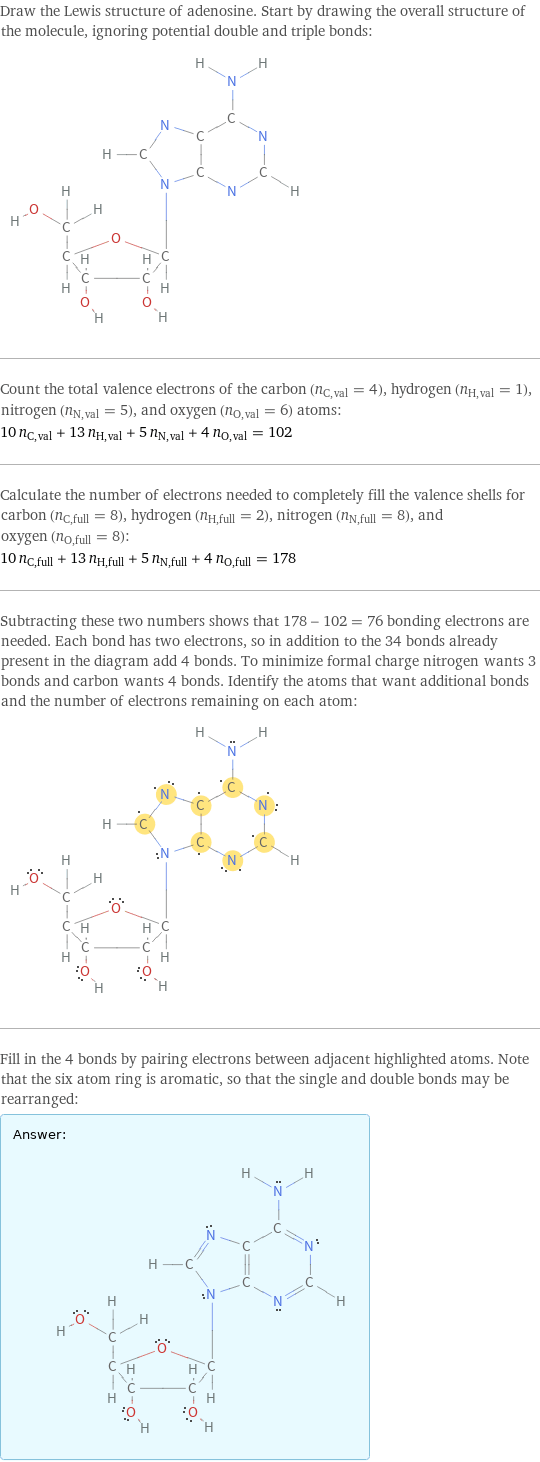
Draw the Lewis structure of adenosine. Start by drawing the overall structure of the molecule, ignoring potential double and triple bonds: Count the total valence electrons of the carbon (n_C, val = 4), hydrogen (n_H, val = 1), nitrogen (n_N, val = 5), and oxygen (n_O, val = 6) atoms: 10 n_C, val + 13 n_H, val + 5 n_N, val + 4 n_O, val = 102 Calculate the number of electrons needed to completely fill the valence shells for carbon (n_C, full = 8), hydrogen (n_H, full = 2), nitrogen (n_N, full = 8), and oxygen (n_O, full = 8): 10 n_C, full + 13 n_H, full + 5 n_N, full + 4 n_O, full = 178 Subtracting these two numbers shows that 178 - 102 = 76 bonding electrons are needed. Each bond has two electrons, so in addition to the 34 bonds already present in the diagram add 4 bonds. To minimize formal charge nitrogen wants 3 bonds and carbon wants 4 bonds. Identify the atoms that want additional bonds and the number of electrons remaining on each atom: Fill in the 4 bonds by pairing electrons between adjacent highlighted atoms. Note that the six atom ring is aromatic, so that the single and double bonds may be rearranged: Answer: | |
3D structure
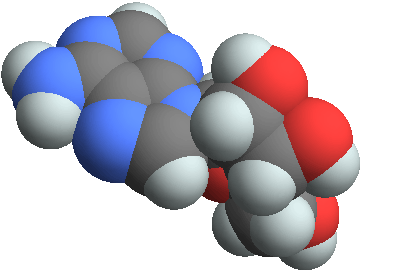
3D structure
Basic properties
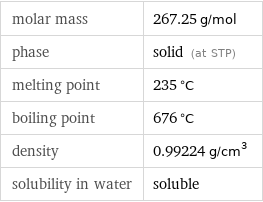
molar mass | 267.25 g/mol phase | solid (at STP) melting point | 235 °C boiling point | 676 °C density | 0.99224 g/cm^3 solubility in water | soluble
Units

Hydrophobicity and permeability properties

experimental LogP hydrophobicity | -1.6 predicted LogP hydrophobicity | -1.2 predicted LogS | -1.28
Drug interactions
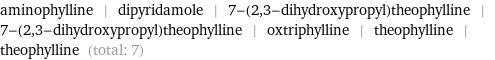
aminophylline | dipyridamole | 7-(2, 3-dihydroxypropyl)theophylline | 7-(2, 3-dihydroxypropyl)theophylline | oxtriphylline | theophylline | theophylline (total: 7)
Basic drug properties
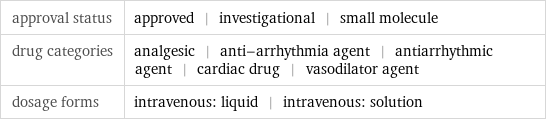
approval status | approved | investigational | small molecule drug categories | analgesic | anti-arrhythmia agent | antiarrhythmic agent | cardiac drug | vasodilator agent dosage forms | intravenous: liquid | intravenous: solution

brand names | adenocard | adenocor | adenoscan | adenosin | adensoine | boniton | myocol | nucleocardyl | pallacor | sandesin
Solid properties (at STP)
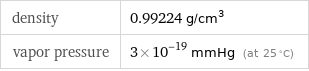
density | 0.99224 g/cm^3 vapor pressure | 3×10^-19 mmHg (at 25 °C)
Units

Thermodynamic properties
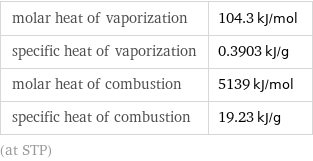
molar heat of vaporization | 104.3 kJ/mol specific heat of vaporization | 0.3903 kJ/g molar heat of combustion | 5139 kJ/mol specific heat of combustion | 19.23 kJ/g (at STP)
Chemical identifiers

CAS number | 58-61-7 Beilstein number | 93029 PubChem CID number | 60961 PubChem SID number | 3512 SMILES identifier | C1=NC2=C(C(=N1)N)N=CN2C3C(C(C(O3)CO)O)O InChI identifier | InChI=1/C10H13N5O4/c11-8-5-9(13-2-12-8)15(3-14-5)10-7(18)6(17)4(1-16)19-10/h2-4, 6-7, 10, 16-18H, 1H2, (H2, 11, 12, 13)/t4-, 6-, 7-, 10-/m1/s1/f/h11H2 InChI key | OIRDTQYFTABQOQ-ONGDMJDYDQ EU number | 200-389-9 RTECS number | AU7175000 NSC number | 7652
NFPA label

NFPA label

NFPA health rating | 1 NFPA fire rating | 0 NFPA reactivity rating | 0
Safety properties

flash point | 362 °C
Toxicity properties

RTECS classes | drug | mutagen | human data